Explaining Predictions of Machine Learning Models with LIME - Münster Data Science Meetup
Slides from Münster Data Science Meetup
These are my slides from the Münster Data Science Meetup on December 12th, 2017.
knitr::include_url("https://shiring.github.io/netlify_images/lime_meetup_slides_wvsh6s.pdf")My sketchnotes were collected from these two podcasts:
- https://twimlai.com/twiml-talk-7-carlos-guestrin-explaining-predictions-machine-learning-models/
- https://dataskeptic.com/blog/episodes/2016/trusting-machine-learning-models-with-lime

Sketchnotes: TWiML Talk #7 with Carlos Guestrin – Explaining the Predictions of Machine Learning Models & Data Skeptic Podcast - Trusting Machine Learning Models with Lime
Example Code
- the following libraries were loaded:
library(tidyverse) # for tidy data analysis
library(farff) # for reading arff file
library(missForest) # for imputing missing values
library(dummies) # for creating dummy variables
library(caret) # for modeling
library(lime) # for explaining predictionsData
The Chronic Kidney Disease dataset was downloaded from UC Irvine’s Machine Learning repository: http://archive.ics.uci.edu/ml/datasets/Chronic_Kidney_Disease
data_file <- file.path("path/to/chronic_kidney_disease_full.arff")- load data with the
farffpackage
data <- readARFF(data_file)Features
- age - age
- bp - blood pressure
- sg - specific gravity
- al - albumin
- su - sugar
- rbc - red blood cells
- pc - pus cell
- pcc - pus cell clumps
- ba - bacteria
- bgr - blood glucose random
- bu - blood urea
- sc - serum creatinine
- sod - sodium
- pot - potassium
- hemo - hemoglobin
- pcv - packed cell volume
- wc - white blood cell count
- rc - red blood cell count
- htn - hypertension
- dm - diabetes mellitus
- cad - coronary artery disease
- appet - appetite
- pe - pedal edema
- ane - anemia
- class - class
Missing data
- impute missing data with Nonparametric Missing Value Imputation using Random Forest (
missForestpackage)
data_imp <- missForest(data)One-hot encoding
- create dummy variables (
caret::dummy.data.frame()) - scale and center
data_imp_final <- data_imp$ximp
data_dummy <- dummy.data.frame(dplyr::select(data_imp_final, -class), sep = "_")
data <- cbind(dplyr::select(data_imp_final, class), scale(data_dummy,
center = apply(data_dummy, 2, min),
scale = apply(data_dummy, 2, max)))Modeling
# training and test set
set.seed(42)
index <- createDataPartition(data$class, p = 0.9, list = FALSE)
train_data <- data[index, ]
test_data <- data[-index, ]
# modeling
model_rf <- caret::train(class ~ .,
data = train_data,
method = "rf", # random forest
trControl = trainControl(method = "repeatedcv",
number = 10,
repeats = 5,
verboseIter = FALSE))model_rf## Random Forest
##
## 360 samples
## 48 predictor
## 2 classes: 'ckd', 'notckd'
##
## No pre-processing
## Resampling: Cross-Validated (10 fold, repeated 5 times)
## Summary of sample sizes: 324, 324, 324, 324, 325, 324, ...
## Resampling results across tuning parameters:
##
## mtry Accuracy Kappa
## 2 0.9922647 0.9838466
## 25 0.9917392 0.9826070
## 48 0.9872930 0.9729881
##
## Accuracy was used to select the optimal model using the largest value.
## The final value used for the model was mtry = 2.# predictions
pred <- data.frame(sample_id = 1:nrow(test_data), predict(model_rf, test_data, type = "prob"), actual = test_data$class) %>%
mutate(prediction = colnames(.)[2:3][apply(.[, 2:3], 1, which.max)], correct = ifelse(actual == prediction, "correct", "wrong"))
confusionMatrix(pred$actual, pred$prediction)## Confusion Matrix and Statistics
##
## Reference
## Prediction ckd notckd
## ckd 23 2
## notckd 0 15
##
## Accuracy : 0.95
## 95% CI : (0.8308, 0.9939)
## No Information Rate : 0.575
## P-Value [Acc > NIR] : 1.113e-07
##
## Kappa : 0.8961
## Mcnemar's Test P-Value : 0.4795
##
## Sensitivity : 1.0000
## Specificity : 0.8824
## Pos Pred Value : 0.9200
## Neg Pred Value : 1.0000
## Prevalence : 0.5750
## Detection Rate : 0.5750
## Detection Prevalence : 0.6250
## Balanced Accuracy : 0.9412
##
## 'Positive' Class : ckd
## LIME
- LIME needs data without response variable
train_x <- dplyr::select(train_data, -class)
test_x <- dplyr::select(test_data, -class)
train_y <- dplyr::select(train_data, class)
test_y <- dplyr::select(test_data, class)- build explainer
explainer <- lime(train_x, model_rf, n_bins = 5, quantile_bins = TRUE)- run
explain()function
explanation_df <- lime::explain(test_x, explainer, n_labels = 1, n_features = 8, n_permutations = 1000, feature_select = "forward_selection")- model reliability
explanation_df %>%
ggplot(aes(x = model_r2, fill = label)) +
geom_density(alpha = 0.5)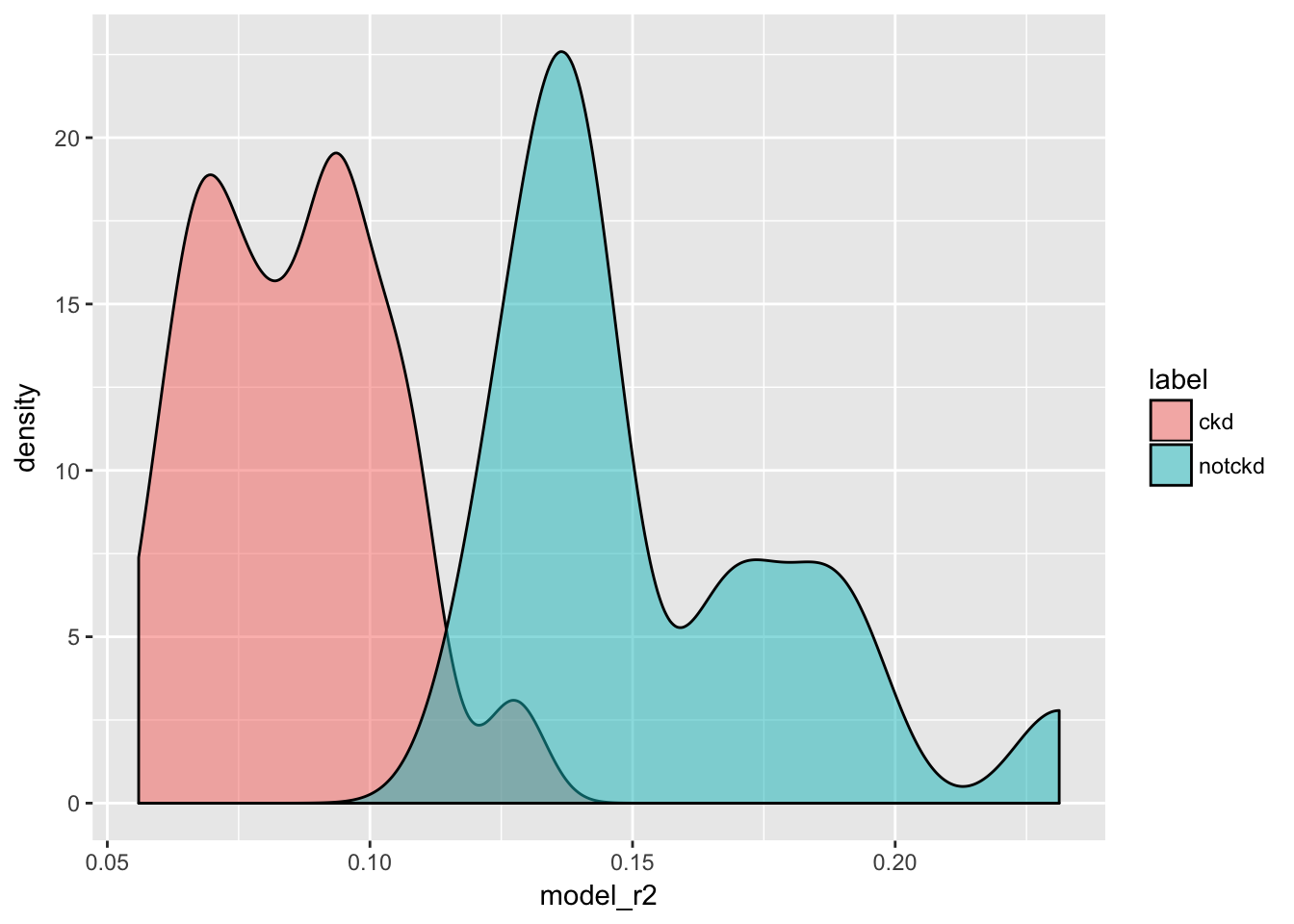
- plot explanations
plot_features(explanation_df[1:24, ], ncol = 1)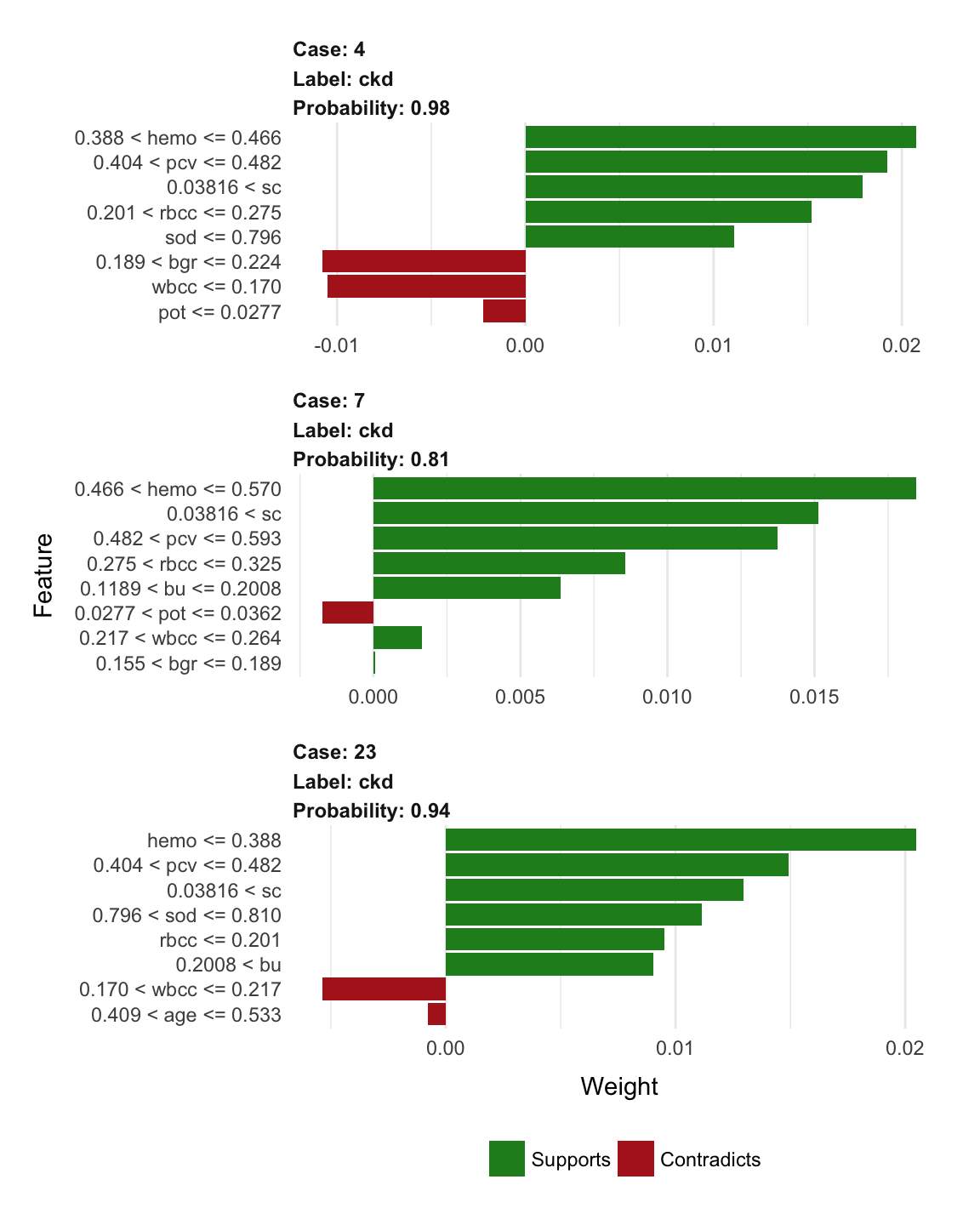
Session Info
## Session info -------------------------------------------------------------## setting value
## version R version 3.4.3 (2017-11-30)
## system x86_64, darwin15.6.0
## ui X11
## language (EN)
## collate de_DE.UTF-8
## tz Europe/Berlin
## date 2018-04-22## Packages -----------------------------------------------------------------## package * version date source
## assertthat 0.2.0 2017-04-11 CRAN (R 3.4.0)
## backports 1.1.2 2017-12-13 CRAN (R 3.4.3)
## base * 3.4.3 2017-12-07 local
## BBmisc 1.11 2017-03-10 CRAN (R 3.4.0)
## bindr 0.1 2016-11-13 CRAN (R 3.4.0)
## bindrcpp * 0.2 2017-06-17 CRAN (R 3.4.0)
## blogdown 0.5 2018-01-24 CRAN (R 3.4.3)
## bookdown 0.7 2018-02-18 CRAN (R 3.4.3)
## broom 0.4.3 2017-11-20 CRAN (R 3.4.2)
## caret * 6.0-78 2017-12-10 CRAN (R 3.4.3)
## cellranger 1.1.0 2016-07-27 CRAN (R 3.4.0)
## checkmate 1.8.5 2017-10-24 CRAN (R 3.4.2)
## class 7.3-14 2015-08-30 CRAN (R 3.4.3)
## cli 1.0.0 2017-11-05 CRAN (R 3.4.2)
## codetools 0.2-15 2016-10-05 CRAN (R 3.4.3)
## colorspace 1.3-2 2016-12-14 CRAN (R 3.4.0)
## compiler 3.4.3 2017-12-07 local
## crayon 1.3.4 2017-09-16 CRAN (R 3.4.1)
## CVST 0.2-1 2013-12-10 CRAN (R 3.4.0)
## datasets * 3.4.3 2017-12-07 local
## ddalpha 1.3.1.1 2018-02-02 CRAN (R 3.4.3)
## DEoptimR 1.0-8 2016-11-19 CRAN (R 3.4.0)
## devtools 1.13.5 2018-02-18 CRAN (R 3.4.3)
## digest 0.6.15 2018-01-28 CRAN (R 3.4.3)
## dimRed 0.1.0 2017-05-04 CRAN (R 3.4.0)
## dplyr * 0.7.4 2017-09-28 CRAN (R 3.4.2)
## DRR 0.0.3 2018-01-06 CRAN (R 3.4.3)
## dummies * 1.5.6 2012-06-14 CRAN (R 3.4.0)
## e1071 1.6-8 2017-02-02 CRAN (R 3.4.0)
## evaluate 0.10.1 2017-06-24 CRAN (R 3.4.1)
## farff * 1.0 2016-09-11 CRAN (R 3.4.0)
## forcats * 0.3.0 2018-02-19 CRAN (R 3.4.3)
## foreach * 1.4.4 2017-12-12 CRAN (R 3.4.3)
## foreign 0.8-69 2017-06-22 CRAN (R 3.4.3)
## ggplot2 * 2.2.1.9000 2018-02-28 Github (thomasp85/ggplot2@7859a29)
## glmnet 2.0-13 2017-09-22 CRAN (R 3.4.2)
## glue 1.2.0 2017-10-29 CRAN (R 3.4.2)
## gower 0.1.2 2017-02-23 CRAN (R 3.4.0)
## graphics * 3.4.3 2017-12-07 local
## grDevices * 3.4.3 2017-12-07 local
## grid 3.4.3 2017-12-07 local
## gtable 0.2.0 2016-02-26 CRAN (R 3.4.0)
## haven 1.1.1 2018-01-18 CRAN (R 3.4.3)
## highr 0.6 2016-05-09 CRAN (R 3.4.0)
## hms 0.4.1 2018-01-24 CRAN (R 3.4.3)
## htmltools 0.3.6 2017-04-28 CRAN (R 3.4.0)
## htmlwidgets 1.0 2018-01-20 CRAN (R 3.4.3)
## httpuv 1.3.6.1 2018-02-28 CRAN (R 3.4.3)
## httr 1.3.1 2017-08-20 CRAN (R 3.4.1)
## ipred 0.9-6 2017-03-01 CRAN (R 3.4.0)
## iterators * 1.0.9 2017-12-12 CRAN (R 3.4.3)
## itertools * 0.1-3 2014-03-12 CRAN (R 3.4.0)
## jsonlite 1.5 2017-06-01 CRAN (R 3.4.0)
## kernlab 0.9-25 2016-10-03 CRAN (R 3.4.0)
## knitr 1.20 2018-02-20 CRAN (R 3.4.3)
## labeling 0.3 2014-08-23 CRAN (R 3.4.0)
## lattice * 0.20-35 2017-03-25 CRAN (R 3.4.3)
## lava 1.6 2018-01-13 CRAN (R 3.4.3)
## lazyeval 0.2.1 2017-10-29 CRAN (R 3.4.2)
## lime * 0.3.1 2017-11-24 CRAN (R 3.4.3)
## lubridate 1.7.3 2018-02-27 CRAN (R 3.4.3)
## magrittr 1.5 2014-11-22 CRAN (R 3.4.0)
## MASS 7.3-49 2018-02-23 CRAN (R 3.4.3)
## Matrix 1.2-12 2017-11-20 CRAN (R 3.4.3)
## memoise 1.1.0 2017-04-21 CRAN (R 3.4.0)
## methods * 3.4.3 2017-12-07 local
## mime 0.5 2016-07-07 CRAN (R 3.4.0)
## missForest * 1.4 2013-12-31 CRAN (R 3.4.0)
## mnormt 1.5-5 2016-10-15 CRAN (R 3.4.0)
## ModelMetrics 1.1.0 2016-08-26 CRAN (R 3.4.0)
## modelr 0.1.1 2017-07-24 CRAN (R 3.4.1)
## munsell 0.4.3 2016-02-13 CRAN (R 3.4.0)
## nlme 3.1-131.1 2018-02-16 CRAN (R 3.4.3)
## nnet 7.3-12 2016-02-02 CRAN (R 3.4.3)
## parallel 3.4.3 2017-12-07 local
## pillar 1.2.1 2018-02-27 CRAN (R 3.4.3)
## pkgconfig 2.0.1 2017-03-21 CRAN (R 3.4.0)
## plyr 1.8.4 2016-06-08 CRAN (R 3.4.0)
## prodlim 1.6.1 2017-03-06 CRAN (R 3.4.0)
## psych 1.7.8 2017-09-09 CRAN (R 3.4.1)
## purrr * 0.2.4 2017-10-18 CRAN (R 3.4.2)
## R6 2.2.2 2017-06-17 CRAN (R 3.4.0)
## randomForest * 4.6-12 2015-10-07 CRAN (R 3.4.0)
## Rcpp 0.12.15 2018-01-20 CRAN (R 3.4.3)
## RcppRoll 0.2.2 2015-04-05 CRAN (R 3.4.0)
## readr * 1.1.1 2017-05-16 CRAN (R 3.4.0)
## readxl 1.0.0 2017-04-18 CRAN (R 3.4.0)
## recipes 0.1.2 2018-01-11 CRAN (R 3.4.3)
## reshape2 1.4.3 2017-12-11 CRAN (R 3.4.3)
## rlang 0.2.0.9000 2018-02-28 Github (tidyverse/rlang@9ea33dd)
## rmarkdown 1.8 2017-11-17 CRAN (R 3.4.2)
## robustbase 0.92-8 2017-11-01 CRAN (R 3.4.2)
## rpart 4.1-13 2018-02-23 CRAN (R 3.4.3)
## rprojroot 1.3-2 2018-01-03 CRAN (R 3.4.3)
## rstudioapi 0.7 2017-09-07 CRAN (R 3.4.1)
## rvest 0.3.2 2016-06-17 CRAN (R 3.4.0)
## scales 0.5.0.9000 2018-02-28 Github (hadley/scales@d767915)
## sfsmisc 1.1-1 2017-06-08 CRAN (R 3.4.0)
## shiny 1.0.5 2017-08-23 CRAN (R 3.4.1)
## shinythemes 1.1.1 2016-10-12 CRAN (R 3.4.0)
## splines 3.4.3 2017-12-07 local
## stats * 3.4.3 2017-12-07 local
## stats4 3.4.3 2017-12-07 local
## stringdist 0.9.4.6 2017-07-31 CRAN (R 3.4.1)
## stringi 1.1.6 2017-11-17 CRAN (R 3.4.2)
## stringr * 1.3.0 2018-02-19 CRAN (R 3.4.3)
## survival 2.41-3 2017-04-04 CRAN (R 3.4.3)
## tibble * 1.4.2 2018-01-22 CRAN (R 3.4.3)
## tidyr * 0.8.0 2018-01-29 CRAN (R 3.4.3)
## tidyselect 0.2.4 2018-02-26 CRAN (R 3.4.3)
## tidyverse * 1.2.1 2017-11-14 CRAN (R 3.4.2)
## timeDate 3043.102 2018-02-21 CRAN (R 3.4.3)
## tools 3.4.3 2017-12-07 local
## utils * 3.4.3 2017-12-07 local
## withr 2.1.1.9000 2018-02-28 Github (jimhester/withr@5d05571)
## xfun 0.1 2018-01-22 CRAN (R 3.4.3)
## xml2 1.2.0 2018-01-24 CRAN (R 3.4.3)
## xtable 1.8-2 2016-02-05 CRAN (R 3.4.0)
## yaml 2.1.17 2018-02-27 CRAN (R 3.4.3)